ByteDance Challenges AlphaFold with Open-Source Protein Prediction Breakthrough
ByteDance Takes on AlphaFold with Open-Source Molecular Modeling Tool
In a move that could democratize computational biology, ByteDance has released Protenix-v1, an open-source model that matches the capabilities of DeepMind's proprietary AlphaFold3 system. This development marks a significant shift in how researchers worldwide might access advanced biomolecular prediction tools.
Breaking Down the Technology Barrier
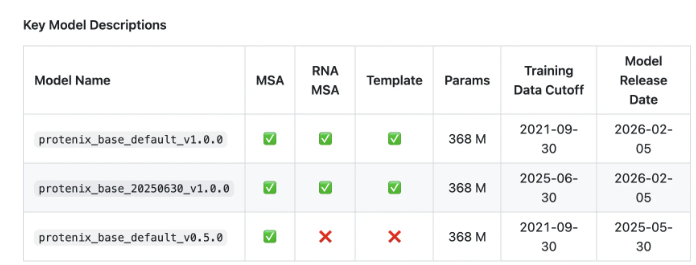
The new model demonstrates remarkable accuracy in predicting full-atom 3D structures across various biological components - from proteins and nucleic acids to small molecule ligands. What makes Protenix-v1 particularly noteworthy is its performance parity with AlphaFold3 despite being freely available under the Apache 2.0 license.
"This isn't just about replicating existing technology," explains Dr. Li Wei, a computational biologist at Peking University. "By open-sourcing both the code and model parameters, ByteDance is removing significant barriers to innovation in fields like drug development."
Setting New Standards for Evaluation
Recognizing the importance of reproducible research, ByteDance simultaneously launched PXMeter v1.0.0, an evaluation toolkit containing over 6,000 complex molecular samples. This comprehensive benchmark allows researchers to validate results and compare different modeling approaches transparently.
The company also introduced a browser-based Web server, enabling scientists without specialized computing resources to conduct interactive research. This accessibility feature could prove particularly valuable for researchers in developing countries or at smaller institutions.
Implications for Scientific Discovery
The open availability of Protenix-v1 could accelerate progress in several cutting-edge areas:
- Drug discovery: Faster identification of potential drug candidates
- Synthetic biology: Improved design of novel biological systems
- Basic research: Enhanced understanding of molecular interactions
While AlphaFold3 remains a powerful tool, the emergence of a comparable open-source alternative may encourage more collaborative development in computational biology. As the field continues to evolve, Protenix-v1 represents both a technological achievement and a philosophical statement about open science.
Key Points:
- ByteDance releases Protenix-v1, an open-source alternative to AlphaFold3
- Model accurately predicts structures for proteins, DNA/RNA, and small molecules
- Includes evaluation toolkit (PXMeter) with 6,000+ molecular samples
- Web server enables browser-based research access
- Potential to accelerate drug discovery and synthetic biology research