MIT and Harvard Unveil Lyra: A Breakthrough in Biological Sequence Modeling
In a significant advancement for computational biology, researchers from MIT, Harvard University, and Carnegie Mellon University have introduced Lyra, a groundbreaking method for biological sequence modeling. This innovation addresses the long-standing challenges of high computational costs and large dataset dependencies in deep learning applications for biology.
A Leap in Efficiency
Lyra represents a paradigm shift in biological sequence analysis, requiring just 1/120,000th of the parameters used in traditional models. Remarkably, the system can be fully trained in two hours using only two GPUs, making advanced biological research accessible to labs with limited resources.
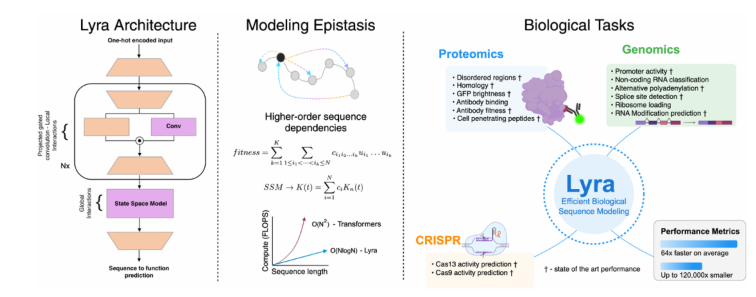
The model's design draws inspiration from epistasis – the interaction between mutations within a sequence. By employing a sub-quadratic architecture, Lyra effectively deciphers the complex relationship between biological sequences and their functions.
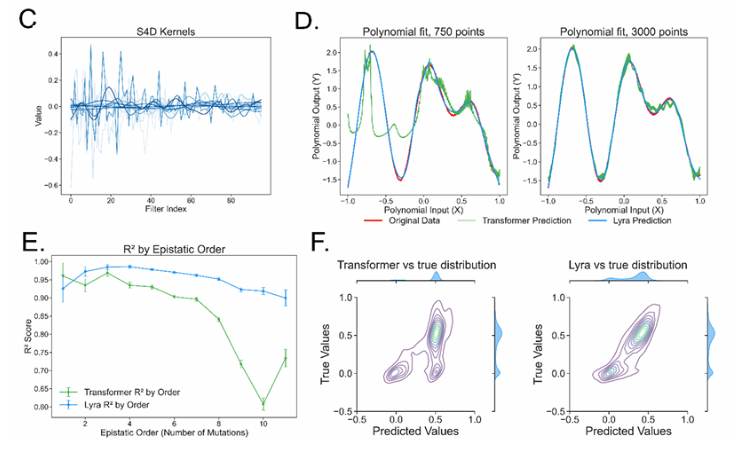
Superior Performance Across Multiple Applications
Testing across over 100 biological tasks, Lyra has demonstrated exceptional results in:
- Protein fitness prediction
- RNA function analysis
- CRISPR design
The system has achieved state-of-the-art (SOTA) performance in several key applications, outperforming conventional methods while using significantly fewer computational resources.
Technical Innovation
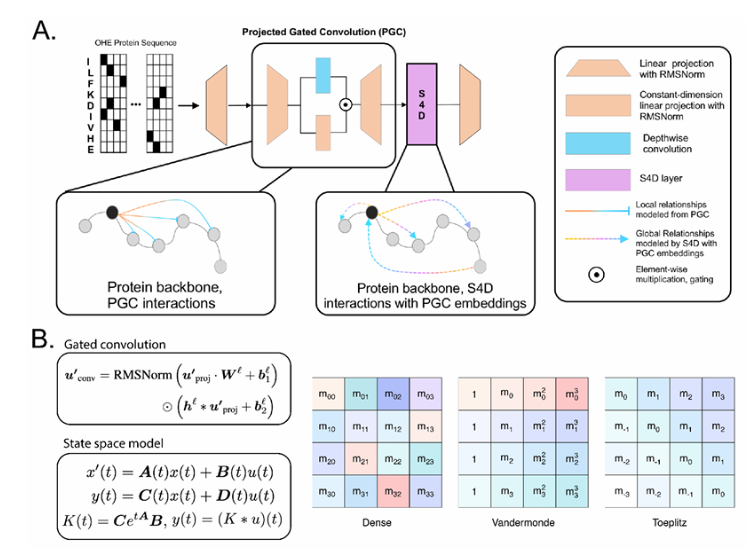
The research team developed a novel hybrid architecture that combines:
- State-space models (SSM) for global relationship modeling using Fast Fourier Transform (FFT)
- Projected gated convolution (PGC) for local feature extraction
This innovative approach results in inference speeds 64.18 times faster than traditional Convolutional Neural Networks (CNNs) and Transformer models.
Broad Implications for Biological Research
The implications of Lyra's efficiency extend across multiple domains:
- Accelerated therapeutic development
- Enhanced pathogen monitoring capabilities
- Improved biomanufacturing processes
The research team hopes their breakthrough will democratize access to advanced biological sequence modeling, enabling more scientists to contribute to cutting-edge discoveries without requiring massive computational infrastructure.
Key Points
- Lyra reduces model parameters by 1/120,000th compared to traditional approaches
- Achieves full training in just two hours using two GPUs
- Demonstrates state-of-the-art performance across 100+ biological tasks
- Features a hybrid SSM-PGC architecture for optimal efficiency
- Runs inferences 64.18× faster than conventional CNNs and Transformers